Identification of Primer Sequences for Genes Associated with Osteoporosis by Eric C. & Kevin L.
Presentation
Summary
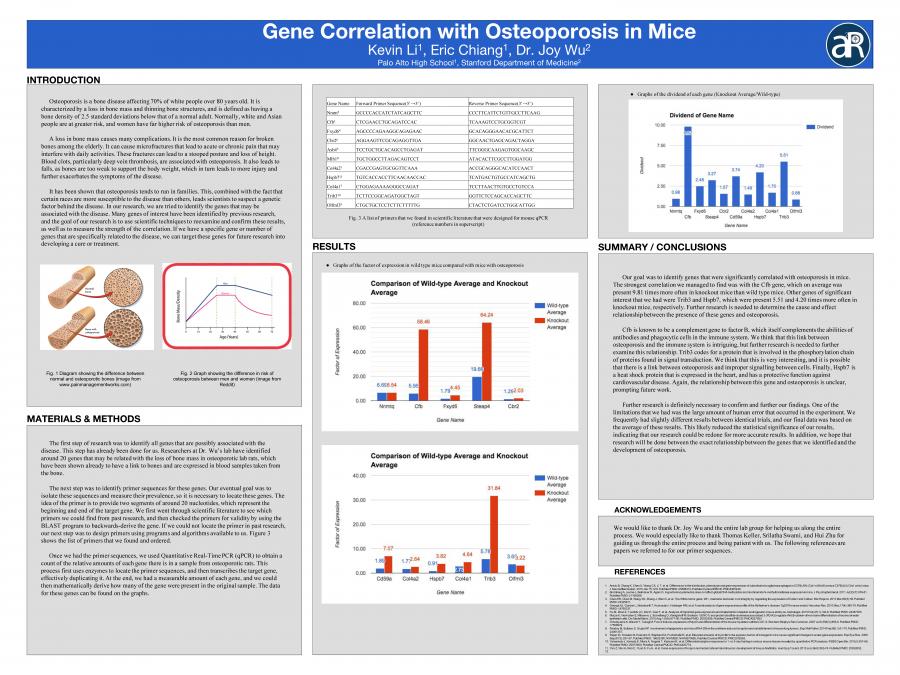
Osteoporosis is a bone disease affecting 70% of white people over 80 years old. It is characterized by a loss in bone mass and thinning bone structures, and is defined as having a bone density of 2.5 standard deviations below that of a normal adult. Normally, white and Asian people are at greater risk, and women have far higher risk of osteoporosis than men. A loss in bone mass causes many complications. It is the most common reason for broken bones among the elderly. It can cause micro-fractures that lead to acute or chronic pain that may interfere with daily activities. These fractures can lead to a stooped posture and loss of height. Blood clots, particularly deep vein thrombosis, are associated with osteoporosis. It also leads to falls, as bones are too weak to support the body weight, which in turn leads to more injury and further exacerbates the symptoms of the disease...It has been shown that osteoporosis tends to run in families. This, combined with the fact that certain races are more susceptible to the disease than others, leads scientists to suspect a genetic factor behind the disease. In our research, we are trying to isolate the genes that may be associated with the disease. If we have a specific gene or number of genes that are specifically related to the disease, we can target these genes for future research into developing a cure or treatment...The first step of research is to identify all genes that are possibly associated with the disease. This step has already been done for us. Researchers at Dr. Wu’s lab have identified around 20 genes that may be related with the loss of bone mass in osteoporotic lab rats, which have been shown already to have a link to bones and are expressed in blood samples taken from the bone. The next step would be to identify primer sequences for these genes. Our eventual goal is to isolate these sequences and measure their prevalence, so this is necessary to locate these genes. The idea of the primer is to provide two segments of around 20 nucleotides, which represent the beginning and end of the target gene. We will first go through scientific literature to see which primers we can find from past research, and then check the primers for validity by using the BLAST program to backwards-derive the gene. If we cannot locate the primer in past research, our next step would be to design primers using programs and algorithms available to us. Once we have the primer sequences, we plan to use Quantitative Real-Time PCR (qPCR) to obtain a count of the relative amounts of each gene there is in a sample from osteoporotic rats. This process first uses enzymes to locate the primer sequences, and then transcribes the target gene, effectively duplicating it. At the end, we have a measurable amount of each gene, and we can then mathematically derive how many of the gene were present in the original sample. We hope to be able to isolate a specific gene or group of genes at this step in order to identify a target for future treatment of the disease.