Summary
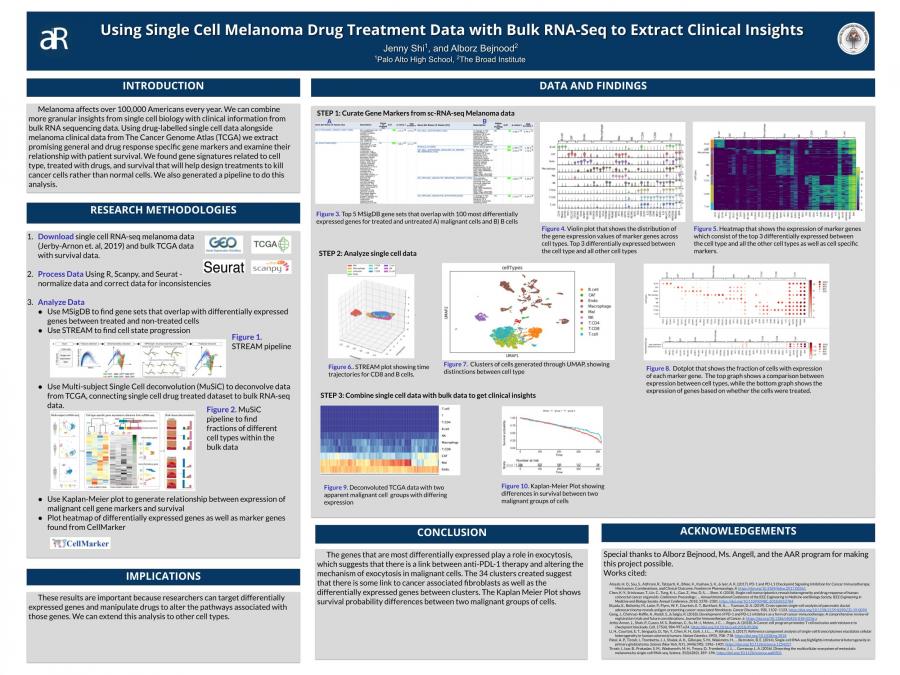
Melanoma affects over 100,000 Americans every year. We can combine more granular insights from single cell biology with clinical information from bulk RNA sequencing data. Using drug-labelled single cell data alongside melanoma clinical data from The Cancer Genome Atlas (TCGA) we extract promising general and drug response specific gene markers and examine their relationship with patient survival. We found gene signatures related to cell type, treated with drugs, and survival that will help design treatments to kill cancer cells rather than normal cells. We also generated a pipeline to do this analysis.
Personal Statement
My project allowed me to pursue my interests in genetics research. After researching nematodes in a project over the summer using RNA sequencing data, I thought it would be interesting to do research using similar data but in a more applicable field of cancer and immunotherapy. AAR has let me read and understand a lot of novel research in this area, as well as develop a lot of new skills in data analysis.